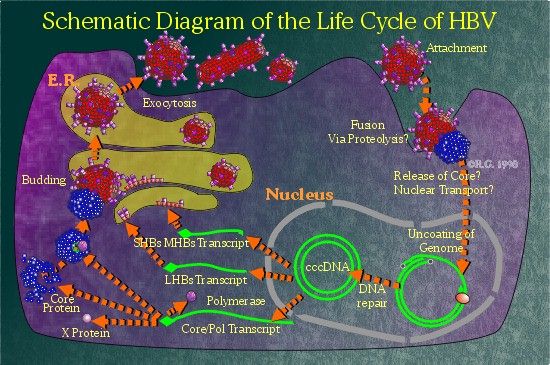
The
Life Cycle of the Hepatitis B Virus
Binding
and Entry
The hepatitis B virus, as in the case of all other viruses, must first attach specifically onto a cell capable of supporting its replication. Though the liver is the most effective cell type for replicating HBV, other extrahepatic sites have been found to be able to support replication to a lesser degree. HBV replicative intermediates and/or viral transcripts have been found in mononuclear cells (1), (2), (3), (4), bile duct epithelial, endothelial, pancreatic acinar cells, and smooth muscle tissue, as well as in adrenal glands, gonads, cultured bone marrow (5), kidneys (6), lymph nodes (7), (8), spleen (9),and thyroid glands of acute hepatitis B infected patients. Viral attachment often determines host and tissue specificity of a virus. However, for HBV, there are no cell-lines available that are able to support viral replication. Only primary duck hepatocytes, which are freshly explanted.from the liver, can support DHBV infection. (10), (11), (12) Consequently, the initial steps of HBV entry is poorly understood. However, several differentiated and immortalized cell lines are capable of supporting viral replication if transfected with viral DNA. (13), (14), (15) The availability of these cell systems has allowed for the elucidation of much of the hepatitis B lifecycle. Many proteins have been found associated with the various hepatitis B surface proteins. Attempts to define the receptor for HBV have yielded a plethora of candidates such as apolipoprotein H(apo-H) (16), an altered form of apolipoprotein H(alt. apo-H) (17), poly-human serum albumin (pHSA) (18), fibronectin (19), and interleukin-6 (IL-6) (20). More recently, a protein (gp180) has been identified to interact with the preS1 domain in DHBV. (21), (22) Also, an 80kd protein has been found to bind onto human HBV which has yet to be identified. (23) Despite these findings, the proteins and mechanism of HBV entry into cells has yet to be well characterized.
Uncoating and Nuclear Transport
The immediate steps following HBV entry are not clearly defined. Some scientists think that the nucleocapsid must be released from the envelope proteins. Other studies suggest that a proteolytic event must occur on the large surface protein to expose a membrane fusion domain. (24) However, it is believed that this occurs at the plasma membrane and not within an acidic vescicle. (25)
After uncoating, the nucleocapsid is believed to be transported to the nuclear membrane. The DHBV system has suggested that HBV genome uncoating occurs at the nuclear membrane. (26)
HBV Genome Repair and Transcription
The HBV DNA is brought into the nucleus where it is repaired to the covalently closed-circular form (cccDNA). Repair requires completion of the dsDNA form, removal of the 5' terminal structures (an RNA primer and the polymerase protein), as well as covalent ligation of the strands. The hepatitis B polymerase protein is not required for these steps to occur. (27) Unlike the retroviridae, integration of HBV DNA into host cell DNA is not required for replication. In fact, integration of HBV DNA results in the disruption of one or more HBV ORFs and prevents transcription of functional pgRNA.
Once recircularized, enhancer and promoter activities start producing the various HBV transcripts required for HBV protein synthesis and pgRNA generation. The transcripts generated can be divided into two categories: subgenomic and genomic. The smaller, subgenomic transcripts serve as mRNA for the expression of the X and surface proteins. The larger genomic transcripts are longer than one genome in length and serve to produce e, core and polymerase proteins. A particular genomic transcript, lacking the ATG start codon for the e protein, is designated the pgRNA since it is chosen specifically by the HBV polymerase protein for packaging. (28) It is hypothesized that ribosome-mediated suppression may account for the inability of HBV to package genomes with the additional e start codon. Mutations which disable this ATG allow the long messages to be encapsidated. (29)
Nucleocapsid Assembly
High levels of core protein expression can lead to empty nucleocapsid formation. (30), (31), (32) However, no genetic information is contained within these empty shells. It is believed that binding of the polymerase protein to the -stem-loop results in pgRNA encapsidation. (33), (34) Deletion of the C-terminus of the polymerase protein allows for binding of the polymerase onto the -stem-loop, but prevents encapsidation. It is believed that the C-terminus of the polymerase is involved in interaction with the hepatitis B core protein. The hepadnaviridae packaging mechanism is different from that found in retroviruses. Retroviruses produce a gag-pol polyprotein which are assembled into functional capsids through interactions between its gag domains. In hepadnaviruses, only polymerase proteins bound into -stem-loops are packaged by core proteins. Expression of core and polymerase proteins alone does not result in the encapsidation of polymerase proteins. (35)
HBV Viral DNA Synthesis
Reverse-transcription of the pgRNA only occurs after core proteins bind to the polymerase. Reverse transcription/DNA replication is believed to be initiated upon commencement of encapsidation.
The complex mechanism employed by HBV to convert its pgRNA into dsDNA is now well-characterized. The transcription of pgRNA results in the generation of a terminally-redundant RNA strand. A sequence, comprised of about 200 nucleotides, including direct repeat 1 (DR1) as well as the -stem-loop, is included. Another copy of DR1 is located near the 3' end of the RNA and denoted as DR2. Upon binding of the polymerase to the -stem-loop, the polymerase begins to reverse transcribe the pgRNA template for three to four bases. The polymerase protein is actually covalently attached onto the growing (-)DNA strand. The polymerase serves as a primer when initiating reverse transcription. Incubating polymerase protein, translated in vitro with mRNA containing the -stem loop and DR1 sequences along with 32P-labeled dNTPs, results in the labeling of the polymerase protein. (36)
The polymerase protein uses a bulge in the -stem-loop as its template for initiating reverse transcription. (37), (38), (39) The four reverse-transcribed nucleotides share identity with four nucleotides in the DR1 region. (40) As such, this segment of DNA can be transferred to the 3' DR1 sequence and reverse transcription can continue by known elongation mechanisms. Reverse transcription of the pgRNA generates the (-)DNA strand which becomes terminally redundant due to the nature of its terminally-redundant template.
As (-)DNA occurs, the newly copied pgRNA template is degraded by RNase H activity of the polymerase protein. (41), (42) However, the 15 to 18 capped oligoribonucleotides at the 5' end of the pgRNA remain undegraded once the (-)DNA strand is completed. This oligoribonucleotide cap serves as the primer for (+)DNA strand synthesis. (43) Generation of this 15 to 18 nucleotide primer is independent of sequence specificity. (44) The newly generated RNA primer is moved and base-paired to the 5' DR2 region on the (-)DNA strand. This step is believed to be facilitated by unidentified proteins. Failure of this translocation event results in the generation of linear dsHBV DNA which occurs in 1% to 5% of primers. (45)
Once translocated, (+)DNA strand synthesis begins and continues to the end of the 5' end of the (-)DNA strand. To complete (+)DNA strand synthesis, an intramolecular transfer is required to give the (+)DNA strand access to the uncopied portion of the (-)DNA strand. This process is facilitated by the terminal-redundancy found in the (-)DNA strand. Typically, the (+)DNA strand is not completed until re-entry of the virion into a host cell. This yields the characteristic single-stranded gap seen in packaged hepadnaviral DNA. One proposed theory is that stearic hindrance within the nucleocapsid prevents further reverse-transcription/replication events. Another suggests that it may be lack of deoxyribonucleic acids in extracellular space which prevents further genome replication. At any rate, incomplete dsDNA/RNA genomes can be readily found in the secreted virions.
Budding and Secretion of Mature Virions and Non-Infectious Particles
Due to the varying amounts of the different surface protein transcripts, much more small and middle hepatitis B surface proteins (SHBs and MHBs) are produced compared to the large hepatitis B surface protein (LHBs). As the S-domain contains several transmembrane spanning sequences, these proteins are co-translated into the endoplasmic reticulum (ER). The hepatitis B surface proteins aggregate in regions of the Golgi complex in a process which excludes host proteins from the complex. Regions high in SHBs protein,with minimal MHBs, are able to bud into the lumen of the ER to produce the secreted 22nm particles. LHBs is rarely contained within 22nm particles as too much LHBs results in ER retention of the particles. (46), (47), (48), (49)
The completed nucleocapsid is thought to associate with areas of the Golgi high in HBV surface protein content. LHBs is believed to interact with the HBc protein at the cytoplasmic face of the Golgi, pulling the nucleocapsid into the forming vesicle, resulting in the 42 nm enveloped particle which is secreted by the cell via exocytosis.
The following section will look more closely at the viral proteins which the HBV genome encodes.
References
1.
Hadchouel M., Pasquinelli, C., Fournier, J.G., et al. 1988. Detection
of Mononuclear Cells Expressing Hepatitis B Virus in Peripheral Blood
from HBsAg Positive and Negative Patients by in situ Hybridization. J.
Med. Virol.; 24: 27-32.
2. Lie-Injo, L.E., Balasegaram, M., Lopez, C.G. and Herrera, A.R. 1983. Hepatitis B Virus DNA in Liver and White Blood Cells of Patients with Hepatoma. DNA; 2: 301-308.
3. Noonan, C.A., Yoffe, B., Mansell, P.W.A., Melnick, J.L. and Hollinger, F.B. 1986. Extrachromosomal Sequences of Hepatitis B Virus DNA in Peripheral Blood Mononuclear Cells of Acquired Immune Deficiency Syndrome Patients. Proc Natl Acad Sci USA; 83: 5698-5702.
4. Shen, H.D., Choo, K.B., Lee, S.D., Tsai, Y.T. and Han, S.H. 1986. Hepatitis B Virus DNA in Leukocytes of Patients with Hepatitis B Virus-Associated Liver Disease. J Med Virol; 18: 201-211.
5. Elfassi, E., Romet-Lemmone, J.L., Essex, M., Frances-McLane, M. and Haseltine, W.A. 1984. Evidence of
Extrachromosomal Forms of Hepatitis B Viral DNA in Bone Marrow Culture Obtained from a Patient Recently Infected with Hepatitis B Virus. Proc Natl Acad Sci USA; 81: 3526-3528.
6. Halpern, M. et al.. 1983. Viral nucleic acid synthesis and antigen accumulation in pancreas and kidneys of Pekin ducks infected with duck hepatitis B virus. Proc. Natl. Acad. Sci. USA; 80: 4865-4869.
7. Laure, F., Zagury, D., Saimot, A.G., Gallo, R.C., Hahn, B.H. and Brechot, C. 1985. Hepatitis B Virus DNA Sequences in Lymphoid Cells from Patients with AIDS and AIDS-related Complex. Science; 229: 561-563.
8. Romet-Lemonne, J.L., McLane, M.F., Elfassi, E., Haseltine, W.A., Azocar, J. and Essex, M. 1983. Hepatitis B Virus Infection in Cultured Human Lymphoblastoid Cells. Science; 221: 667-669.
9. Hosoda, K et al. 1990. Extrahepatic replication of duck hepatitis B virus: more than expected. Hepatology; 11: 44-48.
10. Galle, P.R. et al. 1989. Replication of duck hepatitis B virus in primary duck hepatocytes and its dependence on the state of differentiation of the host cell. Hepatology; 10: 459-465.
11. Pugh, J.C. et al. 1987. Infection and uptake of duck hepatitis B virus by duck hepatocytes maintained in the presence of dimethyl sulfoxide. Virology; 172: 564-572.
12. Tuttleman, J. et al. 1986. In vitro experimental infection of primary duck hepatocytes with duck hepatitis B virus. J. Virol.; 58: 17-25.
13. Sureau, C., Romet-Lemonne, J.L., Mullins, J.I. and Essex, M. 1986. Production of Hepatitis B Virus by a Differentiated Human Hepatoma Cell Line After Transfection with Cloned Circular HBV DNA. Cell; 47: 37-47.
14. Acs, G. et al., 1987. Hepatitis B virus produced by transfected Hep G2 cells causes hepatitis in chimpanzees. Proc. Natl Acad. Sci. USA; 84: 4641-4644.
15. Chang, C. et al. 1987. Production of hepatitis B virus in vitro by transcient expression of cloned HBV DNA in a hepatoma cell line. EMBO J.; 6:675-680.
16. Mehdi, H., Kaplan, M.J., Anlar, F.Y., Yang, X., Bayer, R., Sutherland, K. and Peeples, M.E. 1994. Hepatitis B Virus Surface Antigen Binds to Apolipoprotein H. J Virol; 68: 2415-2424.
17. Mehdi, H., Yang, X. and Peeples, M.E. 1996. An Altered Form of Apolipoprotein H Binds Hepatitis B Virus Surface Antigen Most Efficiently. Virology; 217: 58-66.
18. Pontisso, P, Ruvoletto, M.G., Gerlich, W.H., Heermann, K-H, Bardini, R. and Alberti, A. 1989. Identification of an Attachment Site for Human Liver Plasma Membranes on Hepatitis B Virus Particles. Virology; 173: 522-530.
19. Budkowska, A., Bedossa, P., Groh, F., Louise, A. and Pillot, J. 1995. Fibronectin of Human Liver Sinusoids Binds Hepatitis B Virus: Identification by an Anti-Idiotypic Antibody Bearing the Internal Image of the Pre-S2 Domain. J Virol; 69: 840-848.
20. Neurath, A.R., Strick, N. and Sproul, P. 1992. Search for Hepatitis B Virus Cell Receptors Reveals Binding Sites for Interleukin 6 on the Virus Envelope Protein. J Experimental Medicine; 175: 461-469.
21. 1998. Carboxypeptidase D (gp180), a golgi-resident protein, functions in the attachment and entry of avian hepatitis B viruses - Breiner et al., J. Virol.
22. 1998. Avian hepatitis B virus infection is initiated by the interaction of a distinct pre-S subdomain with the cellular receptor gp180 - Urban et al., J. Virol
23. Ryu, C.J. et al. 2000. An 80-kilodalton protein that binds to the pre-S1 domain of hepatitis B virus. J. Virol.; 74: 110-116.
24. 1996. Protease-induced infectivity of hepatitis B virus for a human hepatoblastoma cell line - Lu et al., J. Virol
25. 1996. Uptake of duck hepatitis B virus into hepatocytes occurs by endocytosis but does not require passage of the virus through an acidic intracellular compartment - Kock et al., J. Virol.
26. 1999. Kinetics of early molecular events in duck hepatitis B virus replication in primary duck hepatocytes - Qiao et al., J. Gen. Virol.
27. Köck, J. et al., 1993. Analysis of the earliest steps of hepadnavirus replication genome repair after infectious entry into hepatocytes does not depend on viral polymerase activity. J. Virol.; 67: 4867-4874.
28. Enders, G.H. et al, 1987. 5' Terminal sequences influence the segregation of ground squirrel hepatitis virus RNAs into polyribosomes and viral core particles. J. Virol.; 61: 35-41.
29. N assal, M. et al., 1990. Translation inactivation of RNA function: discrimination against a subset of genomic transcripts during HBV nucleocapsid assembly. Cell; 63: 1357-1363.
30. Birnbaum, F. et al. 1990. Hepatitis B virus nucleocapsid assembly: primary structure requirements in the core protein. J. Virol.; 64: 3319-3330.
31. Gallina, A. et al., 1989. A recombinant hepatitis B core antigen polypeptide with the protamine-like domain deleted self-assembles into capsid particles but fails to bind nucleic acids. J. Virol; 63: 4645-4652.
32. Miyanohara, A. et al., 1986. Expression of hepatitis B virus core antigen gene in Saccharomyces cerevisiae: syntehsis of two polypeptides translated from different initiation codons. J. Virol.; 1986: 176-180.
33. Hirsch, R. et al. 1990. Polymerase gene products of hepatitis B viruses are required for genomic RNA packaging as well as for reverse transcription. Nature; 344: 552-555.
34. Pollack, J. et al. 1994. Site-specific RNA binding by a hepatitis B virus reverse transcriptase initiated two reactions: RNA packaging and DNA synthesis. J. Virol; 68: 5579-5587.
35. Bartenschlager, R. et al. 1992. Hepadnaviral assembly is initiated by polymerase binding to the encapsidation signal in the viral RNA genome. EMBO J.; 11: 3413-3420.
36. Wang, G-H et al., 1992. The reverse transcriptase of hepatitis B virus acts as a protein primer for viral DNA synthesis. Cell; 71: 663-670.
37. Wang, G-H, et al., 1993. Novel mechanism for reverse transcription in hepatitis B viruses. J. Virol.; 67: 6507-6512.
38. Tavis, J.E. et al. 1993. Expression of functional heaptitis B virus polymerase in yeast reveals it to be the sole viral protein required for correct initiation of reverse transcription. Proc. Natl. Acad. Sci. USA; 90: 4107-4111.
39. Tavis, J.E. et al., 1994. Hepadnaviral reverse transcription initiates within the RNA stem-loop of the viral encapsidation signal and employs a novel strand transfer. J. Virol.; 68: 3536-3543.
40. Wang, G-H et al., 1992. The reverse transcriptase of hepatitis B virus acts as a protein primer for viral DNA synthesis. Cell; 71: 663-670.
41. Miller, R.H. et al., 1984. Hepatitis B DNA-RNA hybrid molecules in particles from infected liver are converted to viral DNA molecules during an endogenous DNA polymerase reaction. Virology; 139: 64-72.
42. Summers, J et al. 1982. Replication of the genome of a hepatitis B-like virus by reverse transcription of an RNA intermediate. Cell; 29: 403-415.
43. Lien, J.M. et al., 1986. Evidence that a capped oligoribonucleotide is the primer for duck hepatitis B plus-strand DNA synthesis. J. Virol.; 57: 229-236.
44. Loeb, D.D. et al., 1991. Sequence-independent RNA cleavages generate the primers for plus strand DNA synthesis in hepatitis B viruses: implications for other reverse transcribing elements. EMBO J; 10: 3533-3540.
45. Staprans, S. et al., 1991. Mutations affecting hepadnavirus plus-strand DNA synthesis dissociate primer cleavage from translocation and reveal the origin of linear viral DNA. J. Virol.;50: 563-571.
46. Cheng, C. et al., 1986. Hepatitis B large surface protein is not secreted but is immunogenic when selectively expressed by recombinant vaccinia virus. J. Virol.; 60: 337-344.
47. Chisari, F. et al. 1986. Expression of hepatitis B large envelope polypeptide inhibits hepatitis B surface antigen secretion in transgenic mice. J. Virol.; 60: 880-887.
48. Persing, D. et al. 1986. Inhibition of secretion of hepatitis B surface antigen by a related presurface polypeptide. Science; 234: 1388-1392.
49. Standring, D.N., 1986. Assembly of viral particles in Xenopus oocytes: pre-surface-antigens regulate secretion of the hepatitis B viral surface envelope particle. Proc. Natl. Acad. Sci. USA; 83: 9338-9342.
